我院张泽教授主持的863子课题“家蚕受驯化和人工选择影响的功能基因组研究(2013AA102507-2)”已经于2017年12月31号顺利结题。
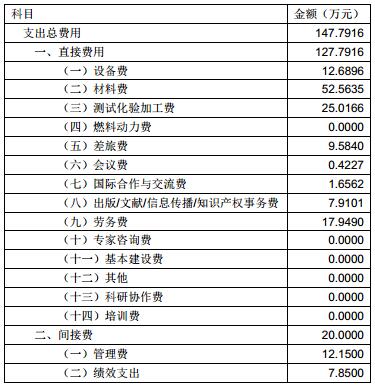
课题组严格按照预算要求使用经费,未进行经费调整。目前共到账经费 190 万元,支出费用147.79 万元。
项目成果概况:
该项目研究了关于家蚕驯化历史及基因流模式分析、家蚕驯化过程中的基因组拷贝数变异、家蚕和野桑蚕丝腺转录组的比较分析、家蚕驯化性状发育整齐度的分子机制、家蚕基因组中大片段重复的鉴定、家蚕孤儿基因的鉴定与进化分析、家蚕Helitron转座子的鉴定和进化分析、家蚕新转座子家族SPY的鉴定和进化分析、转座子的比较基因组学分析、蜕皮激素氧化酶基因的表达调控以及家蚕卵子发生过程中基因表达调控等内容,已在期刊发表论文17篇,达到了项目预期目标。
在该项目的支持下还构建了家蚕转座子数据库和家蚕非编码RNA数据库。
已发表文章:
(1) Zhang Q, Sun W, Sun BY, Xiao Y, Zhang Z* (2017). The dynamic landscapeof gene regulation during Bombyx mori oogenesis. BMC Genomics 18: 71417
(2) Sun W*, Wang CF, Zhang Z (2017). Transcription factor E74A affects the ecdysone titer by regulating the expression of the EO gene in the silkworm,Bombyx mori. BBA-General Subjects 1861:551-558
(3) Zhou QZ, Zhang BD, Yu QY, Zhang Z* (2016). BmncRNAdb: a comprehensive database of non-coding RNAs in the silkworm, Bombyx mori.BMC Bioinformatics. 17:370
(4) Yu QY*, Fang SM, Zhang Z, Jiggins CD* (2016). The transcriptomeresponse of Heliconius melpomene larvae to a novel host plant. Mol Ecol25(19): 4850-65
(5) Tang Z, Zhang HH, Huang K, Zhang XG, Han MJ, Zhang Z* (2015). Repeated horizontal transfers of four DNA transposons in invertebrates andbats. Mobile DNA 6:3
(6) Sun W, Zhao XW and Zhang Z.* (2015). Identification and evolution of theorphan genes in the domestic silkworm, Bombyx mori. Febs Letters589:2731–2738
(7) Fang SM#, Hu BL#, Zhou QZ, Yu QY*, Zhang Z (2015). Comparative analysis of the silk gland transcriptomes between the domestic and wildsilkworms. BMC Genomics 16: 60 (# these authors contributed equally)
(8) Sun W, Shen YH, Han MJ, Cao YF, Zhang Z* (2014). An adaptive transposable element insertion in the regulatory region of the EO gene in thedomesticated silkworm, Bombyx mori. Mol Biol Evol 31: 3302-3313
(9) Yang SY#, Han MJ#, Kang LF, Li ZW, Shen YH*, Zhang Z (2014).Demographic history and gene flow during silkworm domestication. BMCEvol Biol 14:185 (# these authors contributed equally)
(10) Han MJ, Xu HE, Zhang HH, Feschotte C, Zhang Z* (2014). Spy: A new group of eukaryotic DNA transposons without target site duplications.Genome Biol Evol 6(7): 1748-1757
(11) Zhang HH, Feschotte C, Han MJ & Zhang Z* (2014). Recurrent horizontal transfers of Chapaev transposons in diverse invertebrate and vertebrateanimals. Genome Biol Evol 6(6): 1375-1386
(12) Zhao Q, Han MJ, Sun W & Zhang Z* (2014). Copy number variationsamong silkowrms. BMC Genomics 15: 251
(13) Zhang HH, Xu HE, Shen YH, Han MJ & Zhang Z* (2013). The origin and evolution of six miniature inverted-repeat transposable elements (MITEs) in18Bombyx mori and Rhodnius prolixus. Genome Biol Evol 5(11): 2020-2031
(14) Han MJ, Shen YH, Xu MS, Liang HY, Zhang HH & Zhang Z* (2013). Identification and evolution of the silkworm Helitrons and their contributionto transcripts. DNA Res 20: 471-484
(15) Zhao Q, Zhu ZL, Kasahara M, Morishita S & Zhang Z* (2013). Segmentalduplications in the silkworm genome. BMC Genomics 14: 521
(16) Xu HE, Zhang HH, Xia T, Han MJ, Shen YH & Zhang Z* (2013). BmTEdb: a collective database of transposable elements in the silkworm genome.Database (Oxford) 2013: bat055
(17) Zhang HH, Shen YH, Xu HE, Liang HY, Han MJ & Zhang Z* (2013). A novel hAT element in Bombyx mori and Rhodnius prolixus: its relationship with miniature inverted repeat transposable elements (MITEs) and horizontaltransfer. Insect Mol Biol 22(5): 584-596
公示时间:2019.05.10-2019.05.17
生命科学学院
2019.05.10

